|
libSBML C API
5.18.0
|
|
libSBML C API
5.18.0
|
This section presents a complete example program in C++ demonstrating how to create a complete SBML Level 2 model programmatically and then writing the result to a file.
The model presented here is the one described in Section 7.1 of the SBML Level 2 Version 4 specification. It consists of four species identified simply as E, S, ES, and P; one compartment identified as cytosol; and two reactions (one reversible, one irreversible) described by the following equation:
From these equations, it becomes clear that the model also contains three parameters: kon, koff, and kcat.
We will need to assign initial values to these parameters, as well as to the size of the compartment and the initial quantities of the species. Since this model is only meant to illustrate the principles of programming with libSBML, and does not represent an actual biological entity, we hope readers will forgive us for not worrying about justifying the parameter values on the basis of biological plausibility. The table of values for all the quantities we will use is shown below:
| Entity in the Model | Initial value | Units |
|---|---|---|
| cytosol | 1 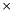 10-14 10-14 | litres |
| S | 1 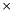 10-20 10-20 | moles |
| P | 0 | moles |
| E | 1 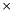 10-21 10-21 | moles |
| ES | 0 | moles |
| kon | 1 000 000 | litre/mole/second |
| koff | 0.2 | 1/second |
| kcat | 0.1 | 1/second |
To add some spice to our boring (but instructive) example, we will use substance units instead of concentration units for the initial values of the species. This is reflected in the use of moles instead of moles/litre as the species quantity units in the table above.
To begin, our program has to include certain header files needed by libSBML under C++. For convenience, we will also set the default C++ namespace to std so that we do not have to prefix common C++ functions used throughout our code.
The next few lines of our program will not be used until the end of the program, but the relevant definitions are best put near the beginning as a principle of good programming. Specifically, this is the definition of a program name and version number, to be inserted by libSBML as XML comments into the final SBML file. We will do something very simple here and define the relevant values as global parameters; other approaches are possible.
We need a few forward declarations for helper functions. Here they are:
Now we can define a main routine for the program. This essentially acts as scaffolding for calling worker functions and dealing with overall errors.
The code above has three key functions, createExampleEnzymaticReaction(), validateExampleSBML, and writeExampleSBML(), which do all the real work in the program. Their verbose names hopefully imply their functions. They are described and defined in the next three subsections.
We begin this function by creating an SBMLDocument_t object, creating a Model_t object within the document object, and setting the Model_t object's identifier.
Next, we need to define some units in order to properly characterize the quantities used in the model. The libSBML API for unit definitions mirrors the SBML object model, which means there is an object class for UnitDefinition and an object class for the Unit objects that can be placed inside a UnitDefinition_t object to compose whatever unit is desired. The resulting code for creating units is a little bit tedious because there is a lot to set up, but on the other hand, it is quite straightforward and hopefully easy to follow:
Before going on, we digress to explain one libSBML API idiom illustrated in the code above and in the rest of the program below. LibSBML provides two main mechanisms for creating objects: class constructors (e.g., Unit_create() ), and createObject() methods provided by certain object classes such as Model_t , where Object is another object class such as UnitDefinition_t . The multiple mechanisms are provided by libSBML for flexibility and to support different use-cases, but they also have different implications for the overall model structure.
In general, the recommended approach is to use the createObject() methods, as in the calls to UnitDefinition::createUnit() above. These methods create an instance and immediately link it into the parent, as well as return a pointer to the object created. By contrast, the other main way of creating an object involves first using the class constructor (e.g., Unit_create() ) to create an object instance, and then calling an addObject(...) method to add the instance to another object. However, the addObject(...) methods in libSBML copy the object instance given to them. The result is that the caller has to be careful about making changes to the object: changes to the original will not affect the copy added to the model by the addObject(...) method, and moreover, the original must be explicitly deleted at some point or else the program will leak memory. In summary, unless special circumstances apply, most applications are best served using the various createObject() methods.
Back to the example now, we create a compartment object:
Next, we turn to the species definitions in the model. For each species, the program needs to set an identifier, the compartment in which it's located, and optionally the initial quantity of the species in the compartment.
It remains only to create objects representing the reactions in the model. In this example, we will create the system as one reversible and one irreversible reaction; alternatively, the system could have been created as three irreversible reactions.
This program has hard-coded versions of the reaction rate formulas only for the purposes of this example. Of course, in a more practical real-life situation, the formulas would not be fixed in this way—one would have methods to read/write/translate expressions originating from different sources (users, files, other programs etc.) to/from the Abstract Syntax Trees (ASTs) used by libSBML.
That was one reaction. One more reaction to go. This time around, for variety and to illustrate another one of libSBML's facilities, we create the AST for the reaction rate formula starting from a string in MathML form.
That concludes all the model creation code. Our createExampleEnzymaticReaction() function now only needs to return the SBMLDocument object that it created.
One of the most important features of libSBML is its ability to perform SBML validation to ensure that a model adheres to the SBML specification for whatever Level+Version combination the model uses. Although the example program presented here is short and we can verify its correctness by inspection, in more general programs, it is difficult to ensure that everything in a model is constructed properly. The ability to perform automatic checks then becomes very useful. We therefore take this opportunity to illustrate the use of libSBML's validation and checking facilities.
The rationale behind libSBML's consistency-checking and validation functions is the following. The individual libSBML for creating and setting attributes and other components are generally quite lenient, and allow a caller to compose SBML entities that may not, in the end, represent valid SBML. This allows applications the freedom to do things such as save incomplete models (which is useful when models are being developed over long periods of time), but at the same time, it means that a separate validation step is necessary when a calling program finally wants to finish a complete SBML document.
LibSBML implements verification of SBML in two steps, represented by the two methods SBMLDocument::checkInternalConsistency() and SBMLDocument::checkConsistency(). The former verifies the basic internal consistency and syntax of an SBML document, and the latter implements more elaborate validation rules (both those defined by the SBML specifications, as well as additional rules offered by libSBML). When an application builds up a model programmatically and is finally ready to say "yes, this model is finished", it should call both of these methods to help ensure the correctness and consistency of the finished result.
The situation is different when an application is reading a model from a file or data stream. Upon reading a model, libSBML automatically verifies the syntax and schema consistency of the document. The application then only needs to check the results by interrogating the error log on the SBMLDocument object and calling SBMLDocument::checkConsistency() to check the model against the SBML validation rules.
The following code illustrates a complete sequence of checking a newly-created model using both SBMLDocument::checkInternalConsistency() and SBMLDocument::checkConsistency(), and handling the reporting of problems to the user.
The final step of writing out a model is relatively trivial. LibSBML provides a class of object (SBMLWriter) that provides the necessary functionality. All that is left to do is to call it with a desired file name.
An expanded version of this example is supplied with the libSBML distribution as the file named createExampleSBML.cpp in the directory examples/c++.